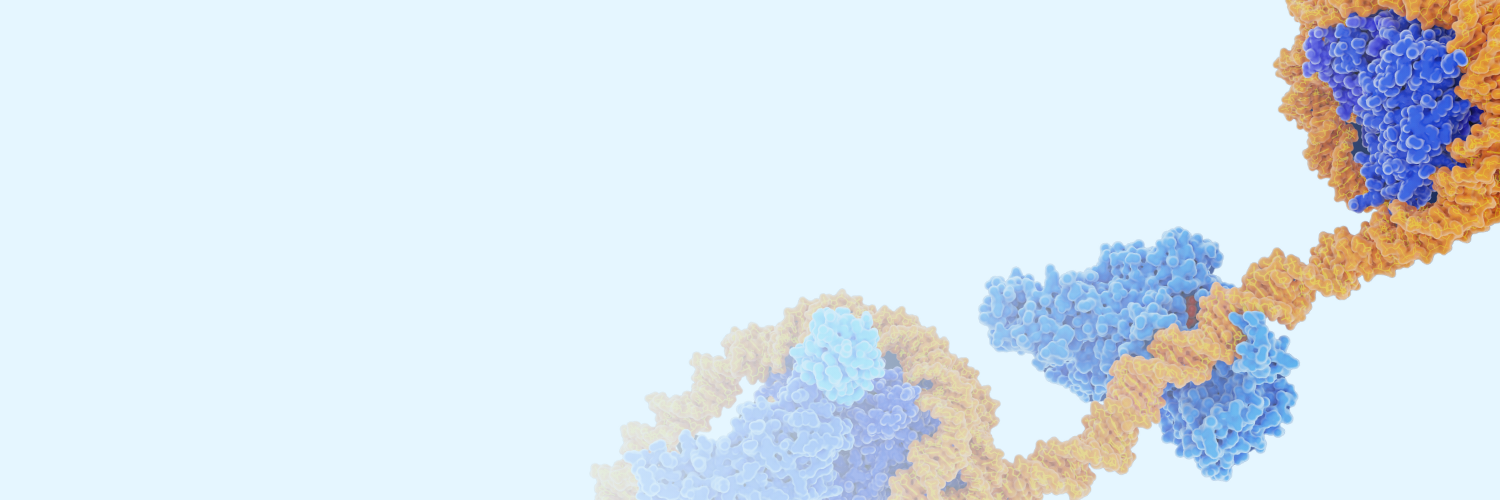

ChIP-seq
Map protein-DNA interactions across the genome to reveal regulatory sites
What is ChIP-seq?
Chromatin immunoprecipitation followed by sequencing (ChIP-seq) is a well-established epigenomic technique used to study DNA-protein interactions across the genome. By using target-specific antibodies to enrich for proteins of interest, such as transcription factors or histone modifications, ChIP-seq enables genome-wide mapping of protein binding sites and chromatin states without prior knowledge of their genomic locations.
ChIP-seq provides valuable insights into gene regulation, chromatin structure, and epigenetic mechanisms, making it a foundational tool for understanding how DNA-protein interactions influence cellular function, development, and disease.

Key Advantages of ChIP-seq

Genome-wide coverage
Map protein-DNA interactions across the entire genome

High resolution & specificity
Identify precise transcription factor binding sites and histone modifications
Versatile
Supports various sample types and input material

Measure relative enrichment to assess changes in protein binding strength between experimental conditions
Quantitative insights
Frequently Asked Questions
Sending samples? See our single-cell sample submission guidelines for best practices on sample preparation, packaging, and shipment for the highest quality results.
-
For ChIP-seq projects, we work with Immunoprecipitated DNA.
Please contact us if you have questions about sample submission guidelines. -
If your sample does not meet our quality control standards at any point in the workflow, we will contact you immediately. We will discuss the specific failure point and provide options, which may include re-submitting the sample, proceeding with a modified workflow, or canceling the project. We believe in transparency and working with you to achieve the best possible outcome.
-
Our workflows are designed to maximize data quality from start to finish. We use specialized protocols for challenging samples like FFPE to ensure high-quality RNA extraction. During library preparation, we use Unique Dual Indexes (UDIs) to minimize index hopping and Unique Molecular Identifiers (UMIs) to correct for PCR duplicates, ensuring accurate quantification. We also perform stringent quality control checks at every step to guarantee reliable and usable data.
-
Bioinformatics for ChIP-seq services include preliminary QC and trimming, genome alignment, duplicate removal, alignment QC, normalization, annotation, enrichment analysis, peak calling & QC, clustering, differential analysis, functional analysis, combined density profile analysis, and motif analysis.
See a ChIP-seq demo report here.
Our team can create custom solutions tailored to your project. Contact us to discuss your project goals and to see how our team can support.
Get the Most out of Your Study
Maximize your project’s potential with advanced analysis solutions. Our team of expert bioinformaticians curate pipelines tailored to your project’s experimental design.
See ChIP-seq in Action
Admera Health provides comprehensive support for all projects, and delivers publication-ready data. Discover how researchers are using Admera Health to advance their studies.
