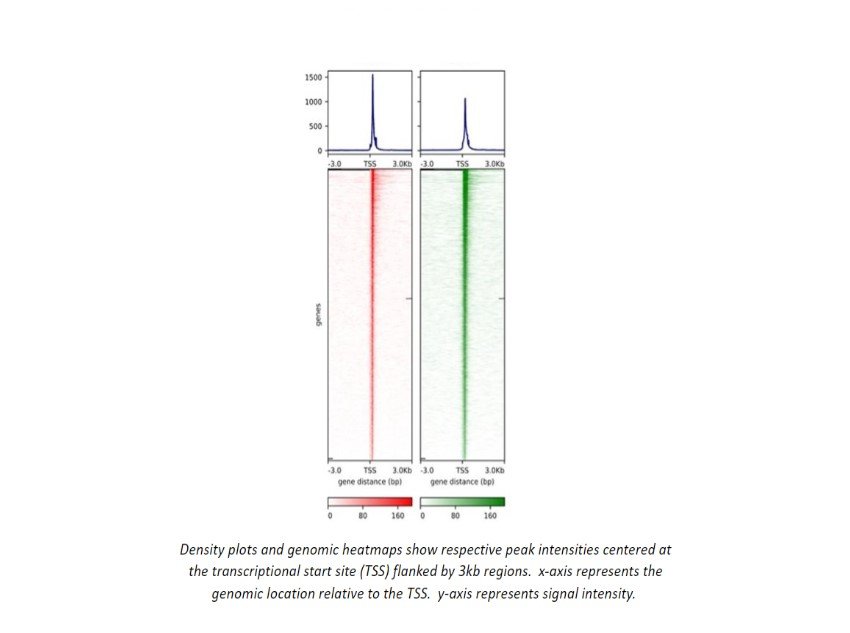
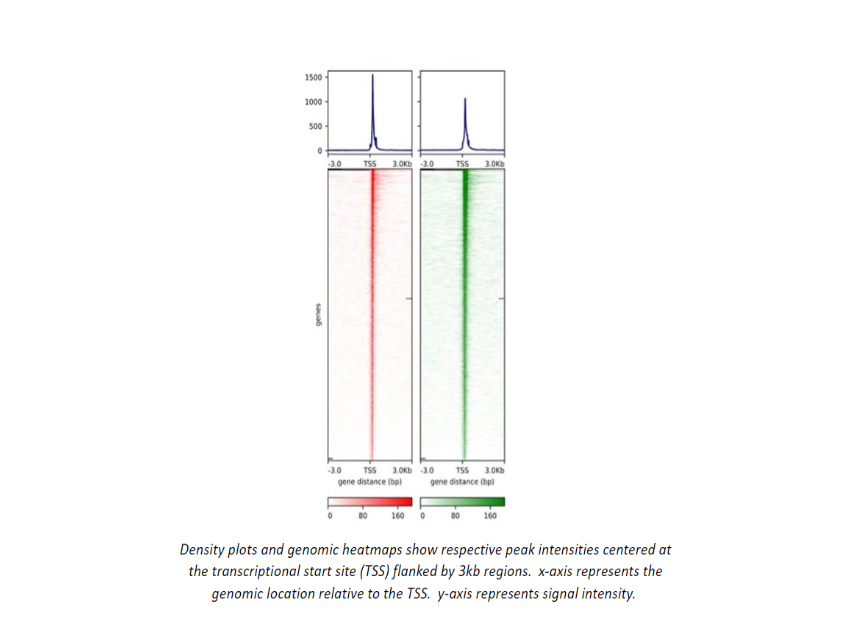
Chip-seq/ATAC-seq/CUT&RUN Demo Data
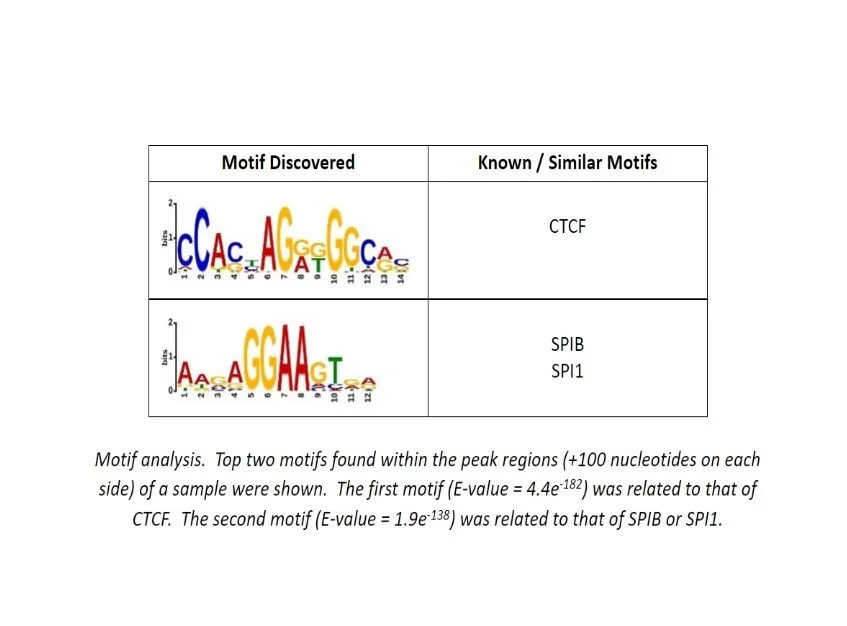
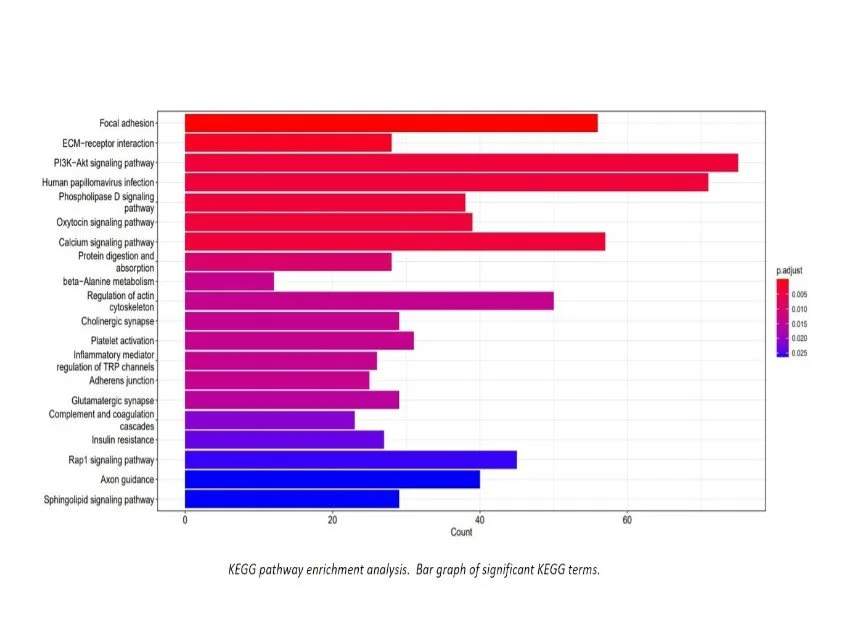
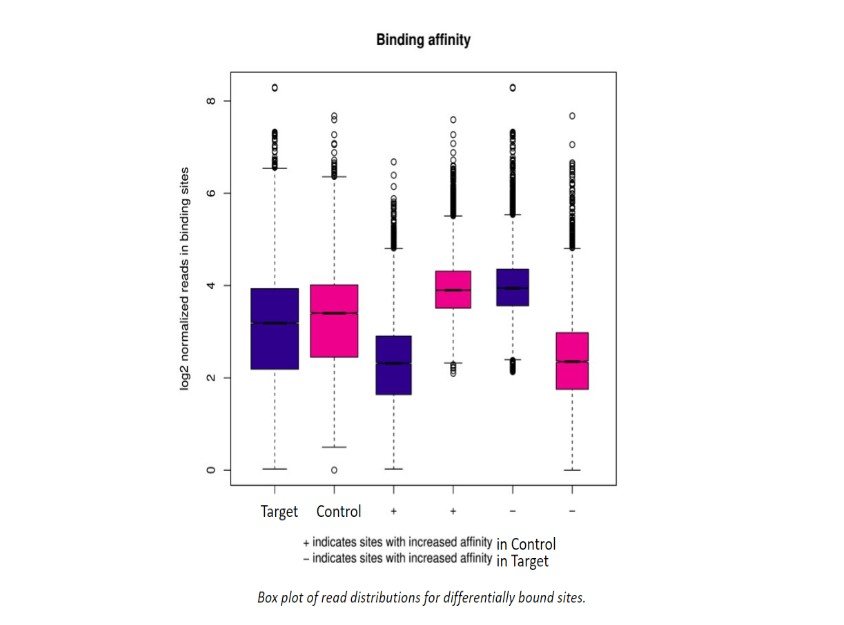
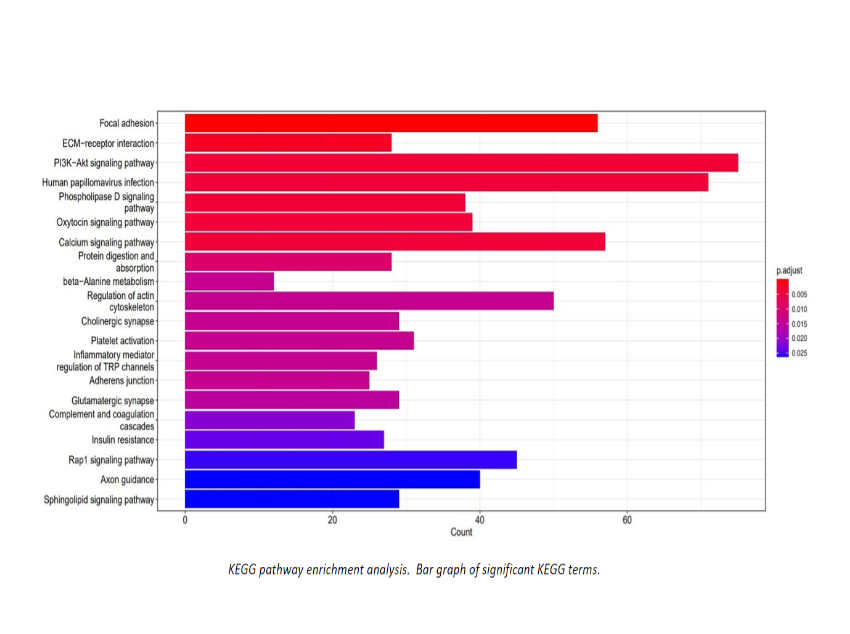
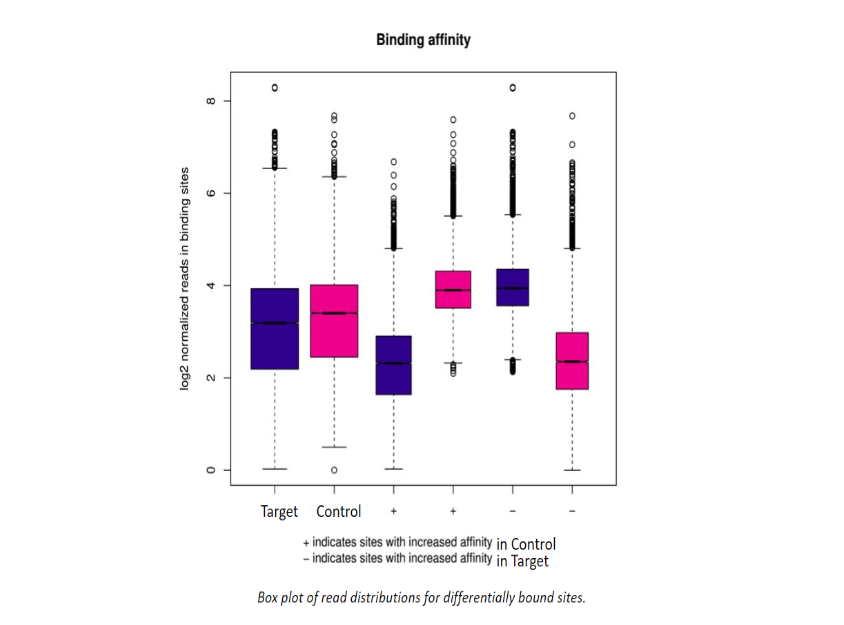
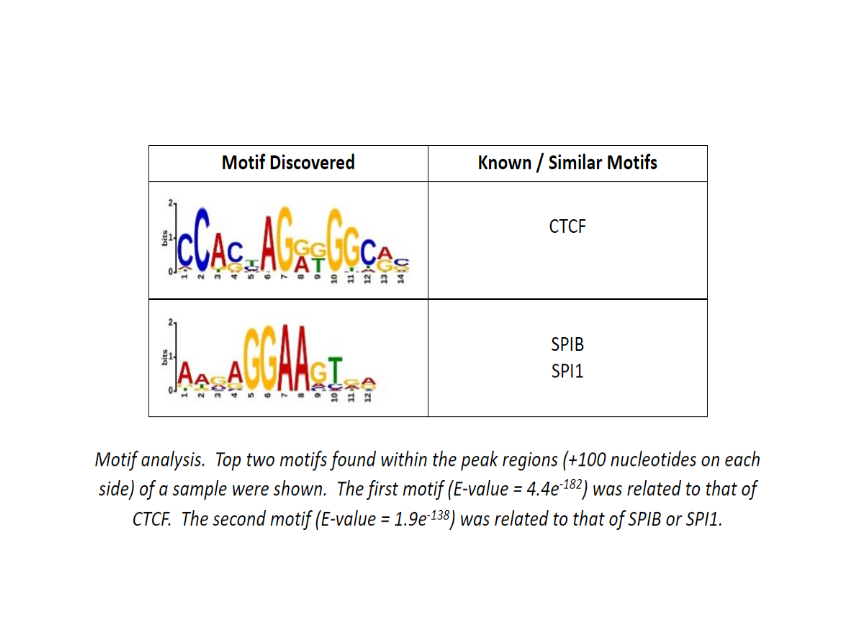
Additional Analysis Figures
Publications
Oh H, Grinberg-Bleyer Y, Liao W, Maloney D, Wang P, Wu Z, Wang J, Bhatt DM, Heise N, Schmid RM, Hayden MS, Klein U, Rabadan R, Ghosh S. An NF-κB Transcription-Factor-Dependent Lineage-Specific Transcriptional Program Promotes Regulatory T Cell Identity and Function. Immunity. 2017 Sep 19;47(3):450-465.e5. doi: 10.1016/j.immuni.2017.08.010. Epub 2017 Sep 7. PMID: 28889947; PMCID: PMC5679261. Library preparation and sequencing were performed by Admera Health. Library was constructed on immunoprecipitated DNA according to manufacturer’s recommendations (KAPA Hyper Prep Kit).
Library preparation and sequencing were performed by Columbia University’s JP Sulzberger Genome Center and Admera Health.
The libraries were quantified using a KAPA Library Quantification kit and sequenced on Illumina HiSeq2000 in paired read mode with the read length of 150 nucleotides at the Admera Health.
DNA fragments were purified from the immunoprecipitated samples, and the corresponding control samples (Input; digested chromatin not subjected to immunoprecipitation) were sequenced on the Illumina platform (Admera Health, NJ, USA) in paired-end, 150 bp mode
Next-generation sequencing (NGS) library preparation and sequencing of one genomic DNA sample, one Input, and one ChIP sample (the first biological replicate using anti-Cg-CenH3) were performed by Admera Health facility (USA) on the Illumina HiSeqX platform, using multiplexing of the samples in a single lane.
The libraries were quantified using a KAPA Library Quantification kit and sequenced on Illumina HiSeq X Five/Ten sequencing systems in paired-end 150bp reads at the Admera Health.
The libraries were sequenced on Illumina’s HiSeq4000 in paired read mode with the read length of 150 nucleotides at the Admera Health.